Abstract
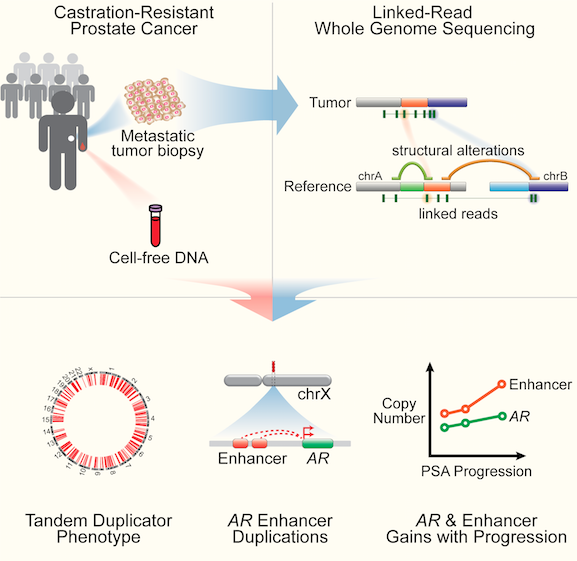
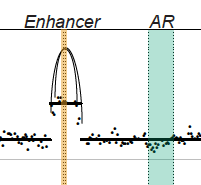
Nearly all prostate cancer deaths are from metastatic castration-resistant prostate cancer (mCRPC), but there have been few whole-genome sequencing (WGS) studies of this disease state. We performed linked-read WGS on 23 mCRPC biopsy specimens and analyzed cell-free DNA sequencing data from 86 patients with mCRPC. In addition to frequent rearrangements affecting known prostate cancer genes, we observed complex rearrangements of the AR locus in most cases. Unexpectedly, these rearrangements include highly recurrent tandem duplications involving an upstream enhancer of AR in 70%–87% of cases compared with <2% of primary prostate cancers. A subset of cases displayed AR or MYC enhancer duplication in the context of a genome-wide tandem duplicator phenotype associated with CDK12 inactivation. Our findings highlight the complex genomic structure of mCRPC, nominate alterations that may inform prostate cancer treatment, and suggest that additional recurrent events in the non-coding mCRPC genome remain to be discovered.